Disentangling primer interactions improves SARS-CoV-2 genome sequencing by multiplex tiling PCR | PLOS ONE
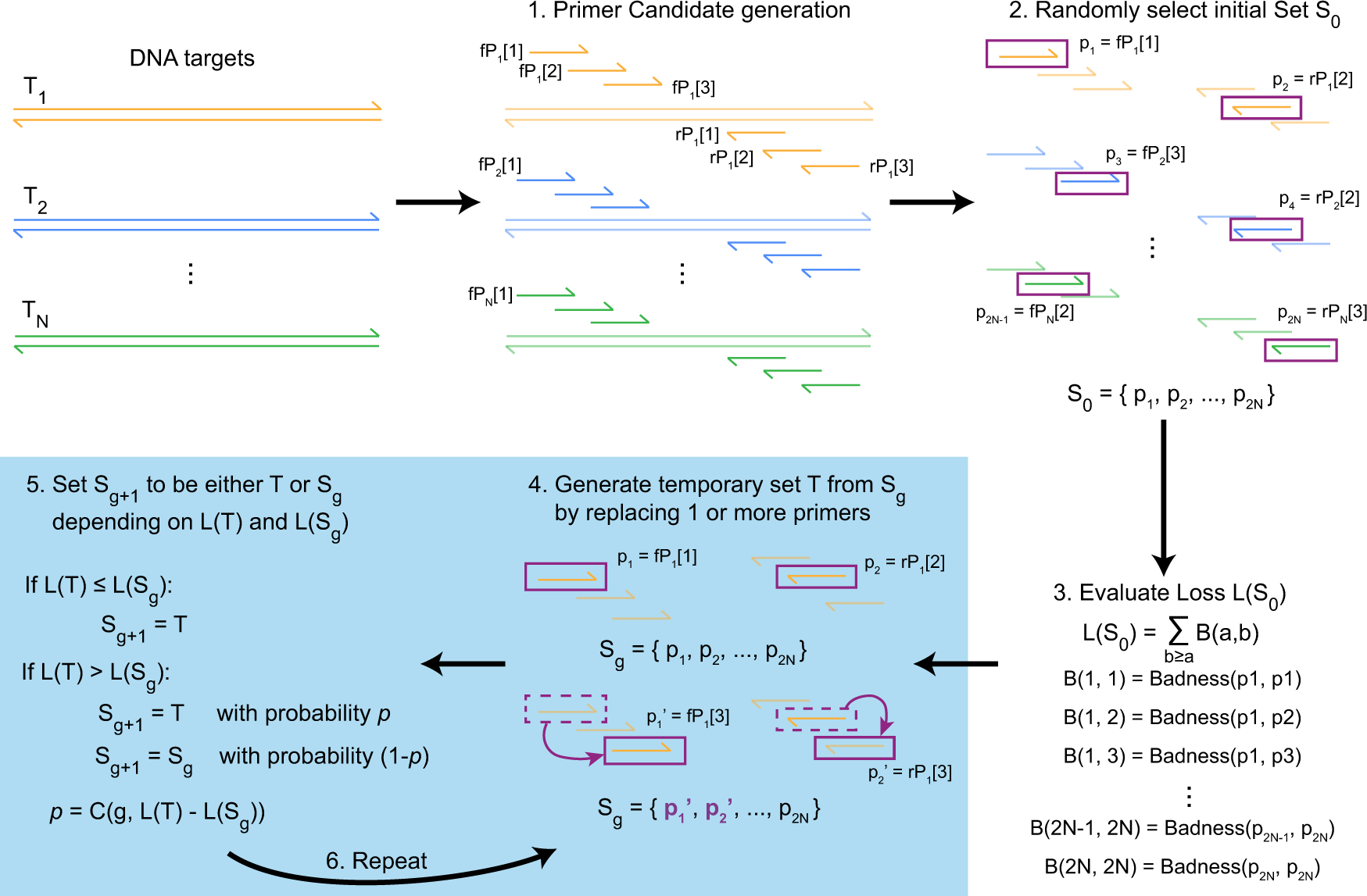
Designing highly multiplex PCR primer sets with Simulated Annealing Design using Dimer Likelihood Estimation (SADDLE) | Nature Communications
The Use of Coded PCR Primers Enables High-Throughput Sequencing of Multiple Homolog Amplification Products by 454 Parallel Sequencing | PLOS ONE

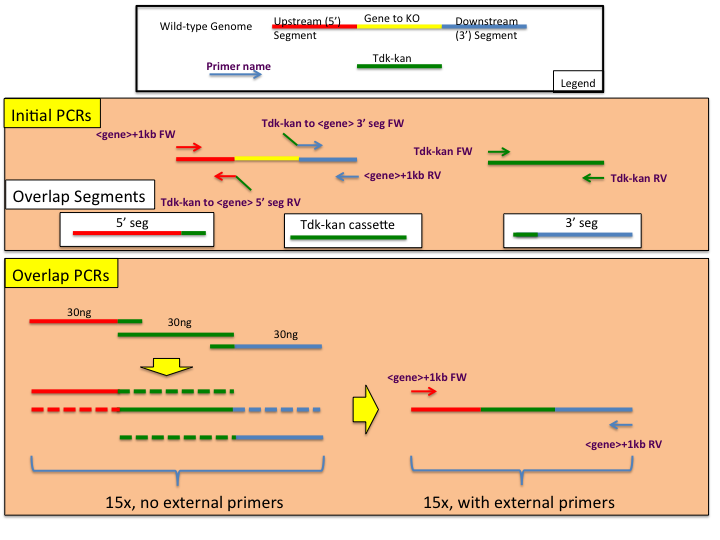
Rapid, accurate mapping of transgene integration in viable rhesus macaque embryos using enhanced-specificity tagmentation-assisted PCR: Molecular Therapy - Methods & Clinical Development
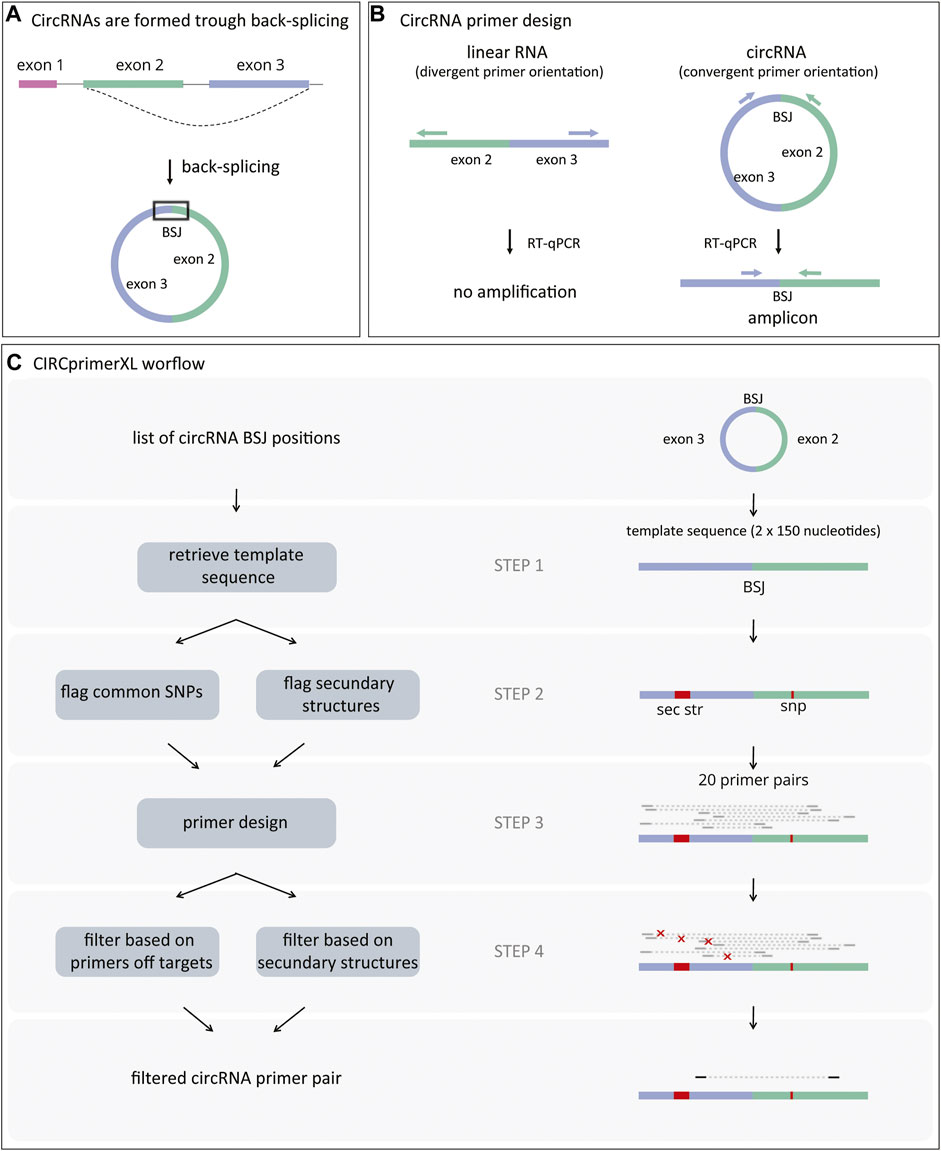
Frontiers | CIRCprimerXL: Convenient and High-Throughput PCR Primer Design for Circular RNA Quantification
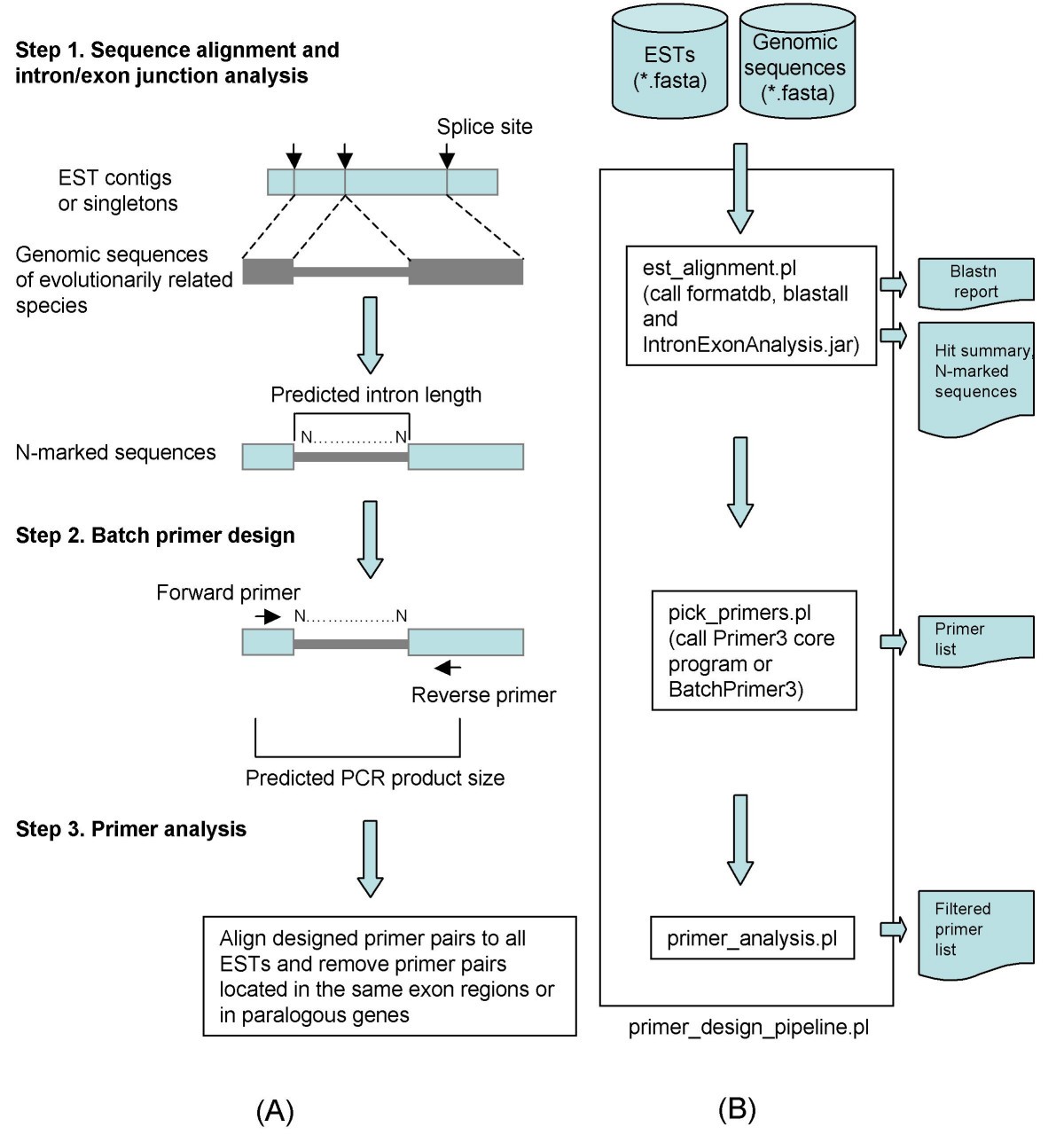
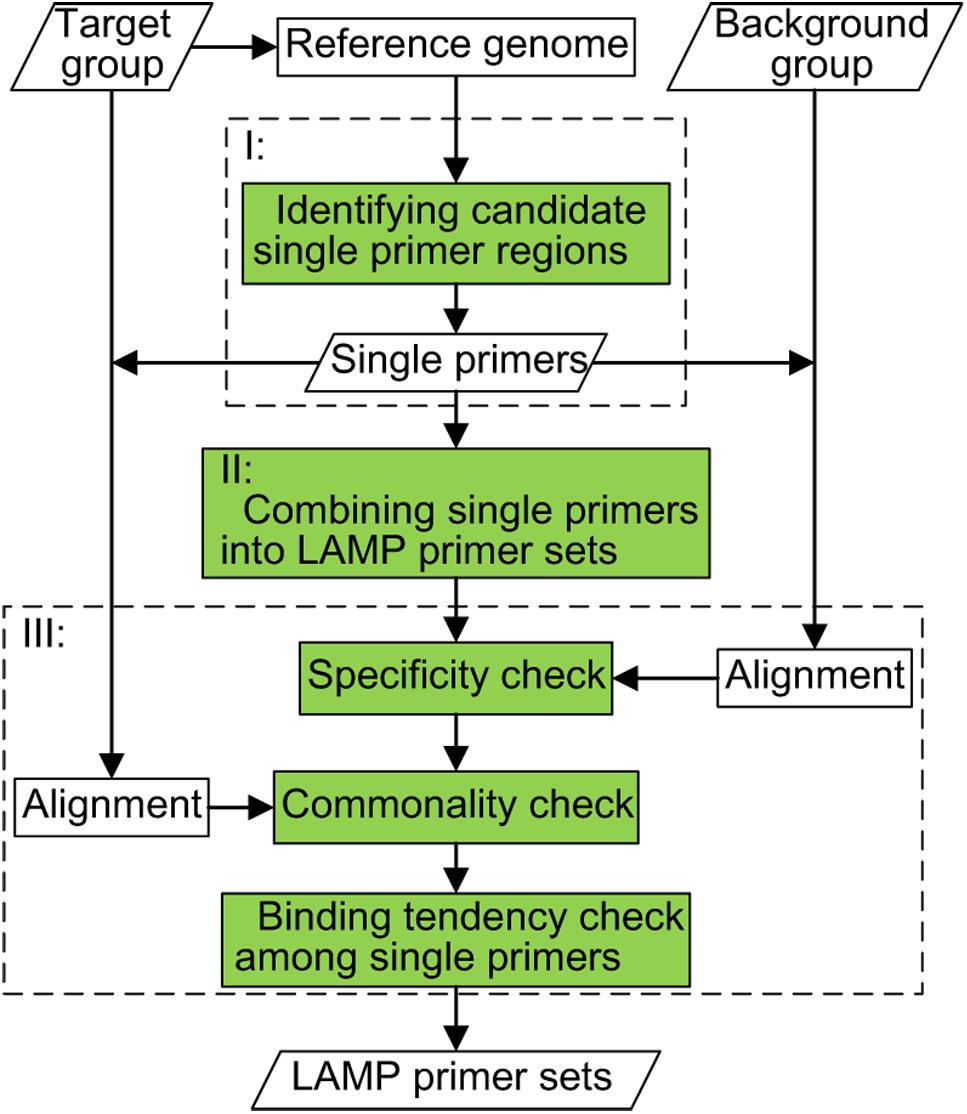
ConservedPrimers 2.0: A high-throughput pipeline for comparative genome referenced intron-flanking PCR primer design and its application in wheat SNP discovery | BMC Bioinformatics | Full Text
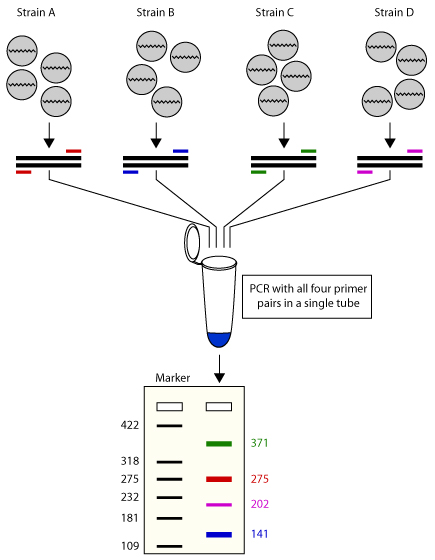
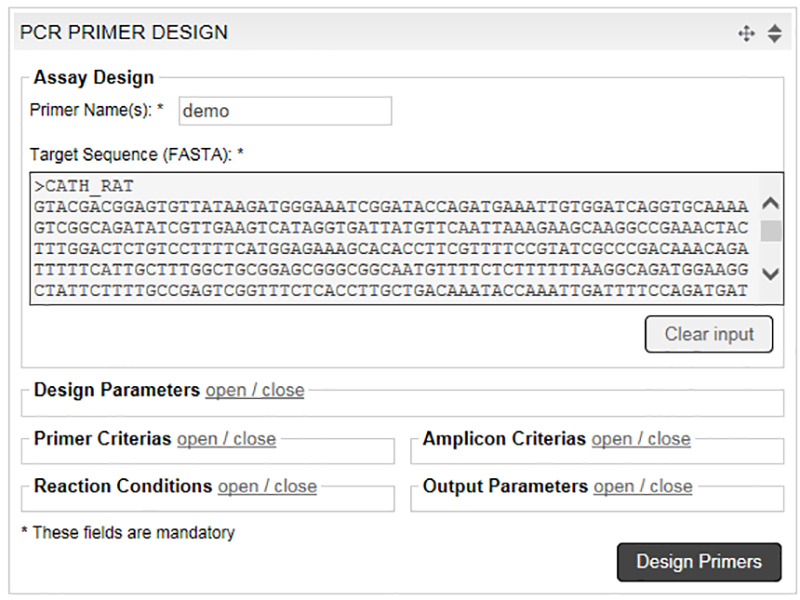
Multiplex PCR: An overview of multiplex PCR assay, primer design for multiplexing, primer design software for multiplex PCR.
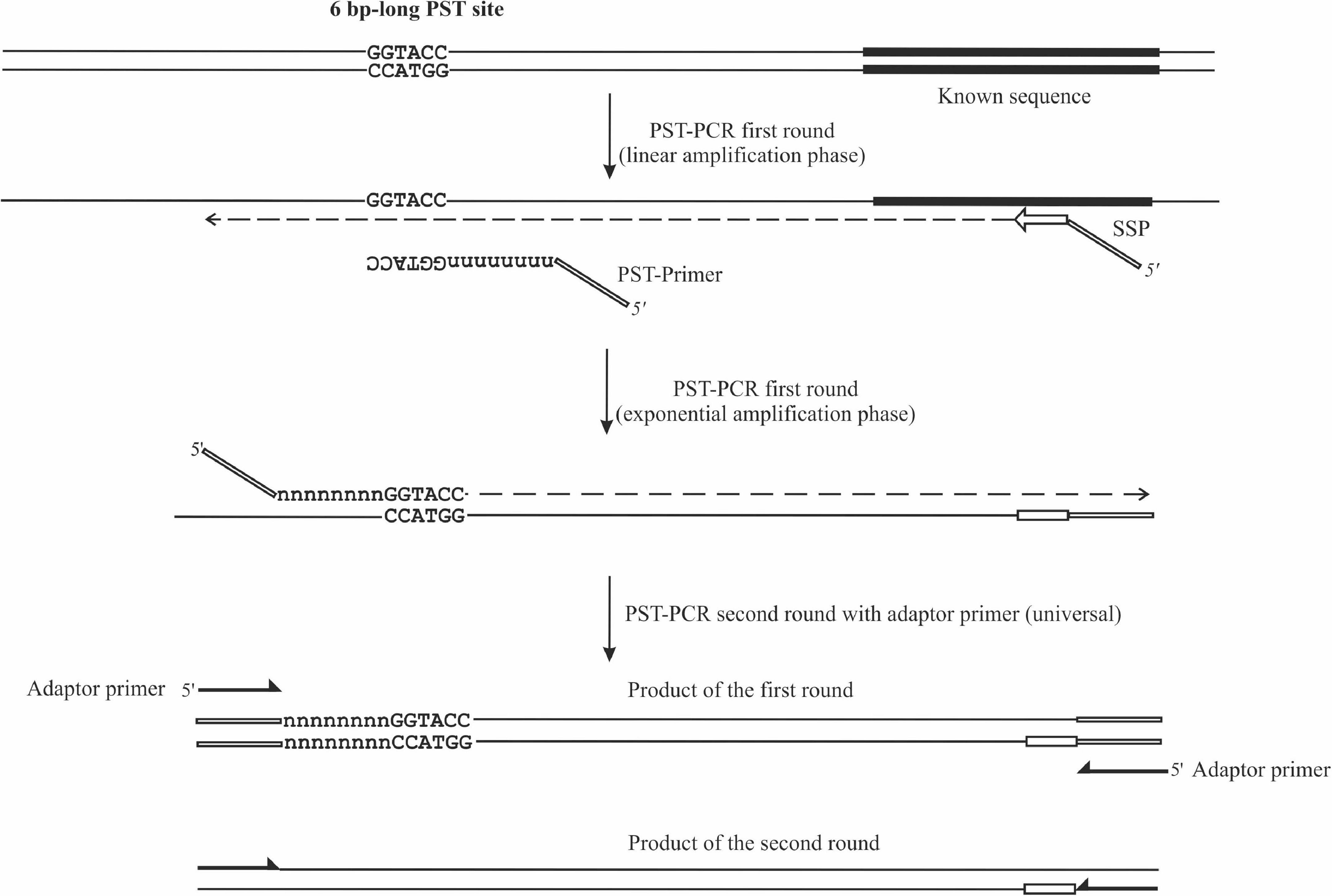
Frontiers | Palindromic Sequence-Targeted (PST) PCR, Version 2: An Advanced Method for High-Throughput Targeted Gene Characterization and Transposon Display